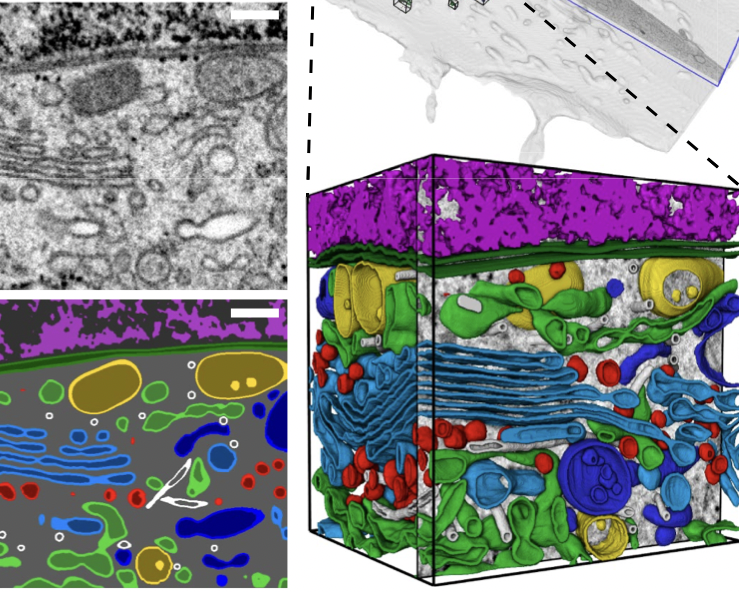
With the help of a convolutional neural network, scientists for the first time managed to carry out a complete 3D reconstruction of a biological cell based on electron microscopy data. The reconstruction process using a neural network takes several hours, which is about 100,000 times faster than with manual data processing.
Cells consist of many organelles that control their activity. Despite the fact that these structures are well studied, there has not yet been a map of their spatial distribution on a nanometer scale. Such a map can be obtained using ultra-high resolution electron microscopy techniques. However, the data for only one cell consists of tens of thousands of images, and therefore the analysis of all cell organelles would take one scientist more than 60 years.
The team of scientists managed to solve this problem using the 3D U-net neural network, which classified thousands of organelles inside the cell in a few hours. The neural network was trained on 30 manually marked types of organelles and other structures.
The key result of the study is to obtain information about how much space organelles occupy inside the cell, how they are located relative to each other and how they interact. In addition to two articles in Nature, scientists have launched the OpenOrganelle portal, where arrays of neural network output data, datasets and tools for processing electron microscopy data are publicly available.


