Researchers at the Massachusetts Institute of Technology used a convolutional neural network to search for highly active and non-toxic peptides. The neural network proposed previously unknown peptides that allowed to increase the delivery rate of the drug for the treatment of muscular dystrophy by 50 times and reduce its toxicity.
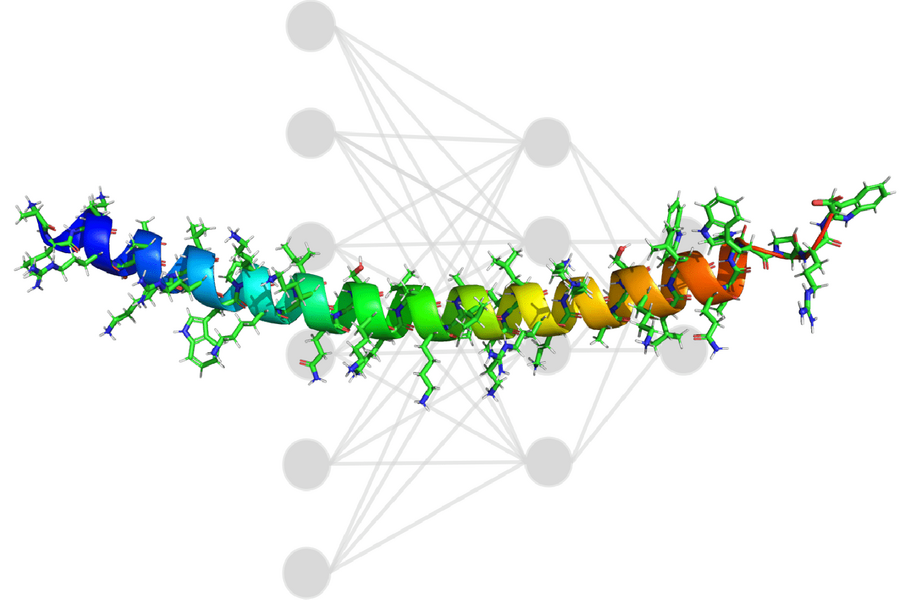
The drug for the treatment of Duchenne muscular dystrophy (DMD) uses large synthetic PMO molecules that penetrate the cell nucleus and modify the dystrophin gene in such a way as to start the production of proteins that are absent in patients with DMD. The key problem of PMO molecules is their low ability to penetrate into cells. To speed up the delivery of PMO to the nucleus, pharmacists attach peptides that penetrate into cells to the molecule, thereby helping it cross cell membranes and reach the target. However, it was previously unclear how to solve the problem of determining the most suitable sequence of peptides: for example, the number of peptides with 40 amino acid residues exceeds the number of atoms on Earth.
MIT researchers were able to use AI to detect non-toxic and highly active peptides that can be attached to PMO to facilitate delivery. The results of the study are published in Nature Chemistry. To collect the training dataset, a group of scientists experimentally compiled a set of 600 peptide sequences, each of which was attached to a PMO, and quantified the activity (delivery efficiency) of each variant.
After training, the model learned to predict the activity of peptide sequences and proposed sequences that turned out to be more effective than any previously known variants. One of them, in particular, can increase the delivery of PMO by 50 times compared to the delivery speed of the current version of the drug. The researchers injected a modified version of the drug into mice and confirmed the prognosis of the neural network, as well as demonstrated the non-toxicity of the new sequences.
The source code of the model was posted on Github, and third-party scientists have already used it to improve and develop more than 10 other types of drugs.