MedSAM2: Open Source SOTA 3D Medical Image and Video Segmentation Model
13 April 2025
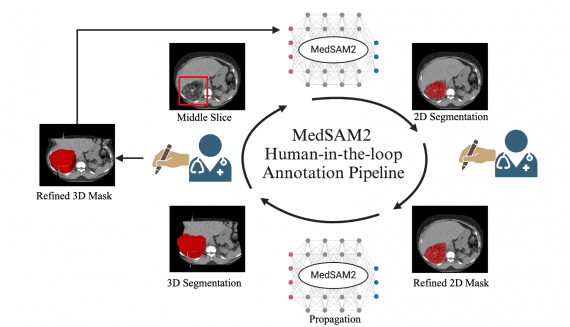
MedSAM2: Open Source SOTA 3D Medical Image and Video Segmentation Model
Medical image segmentation plays a critical role in precision medicine, enabling more accurate diagnosis, treatment planning, and quantitative analysis. While significant progress has been made in developing both specialized and…